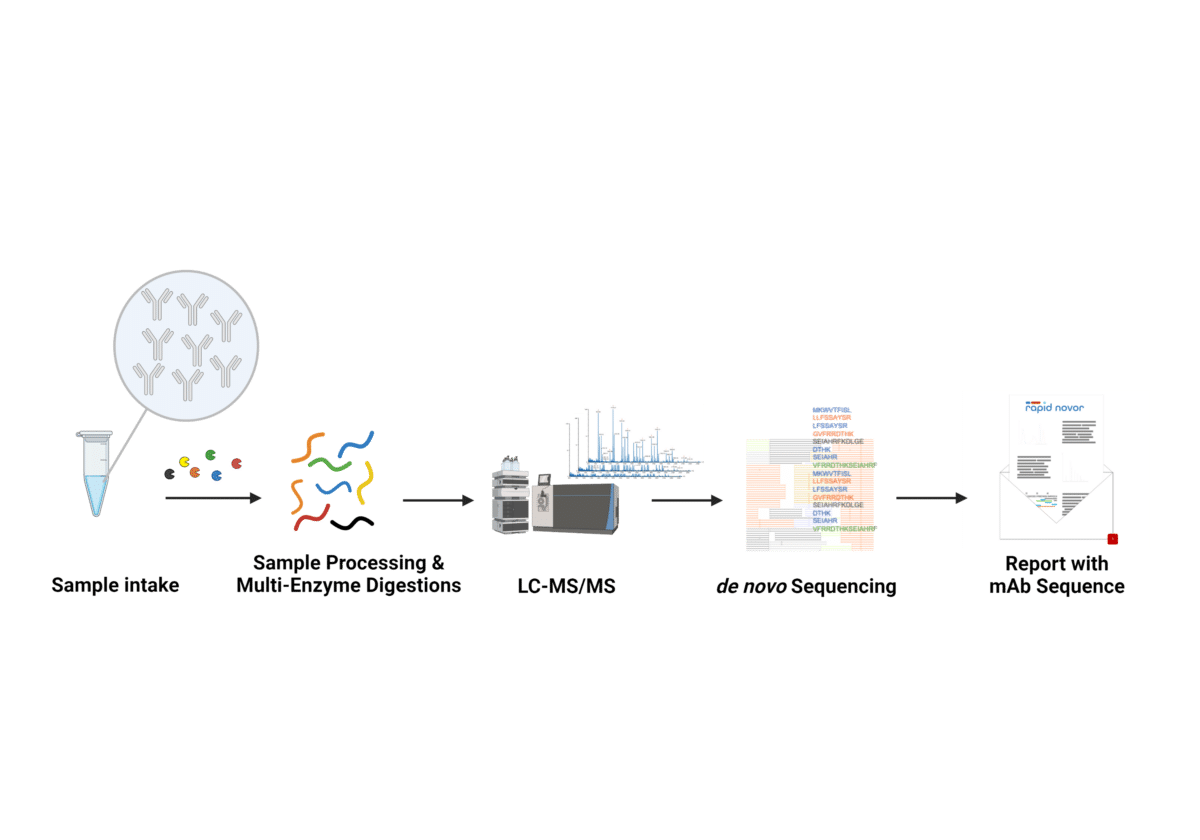
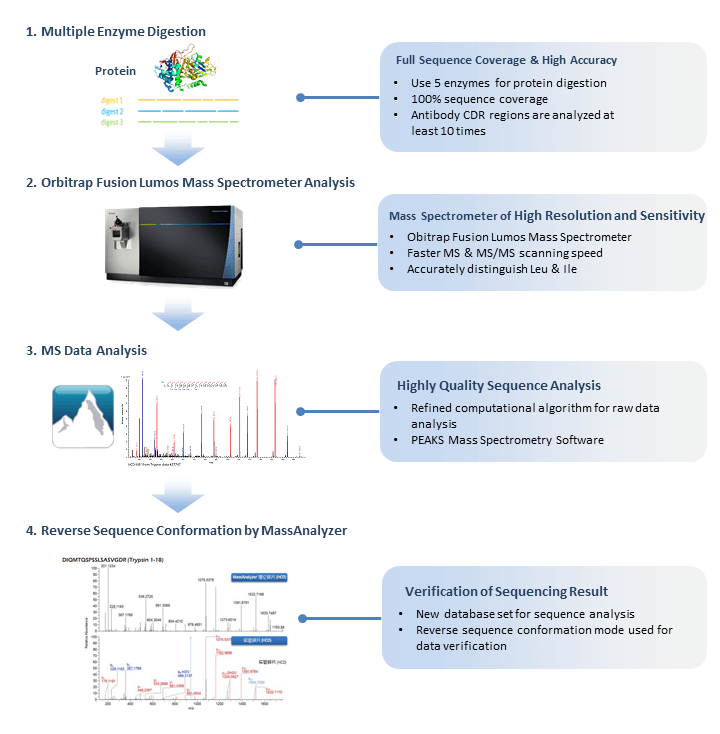
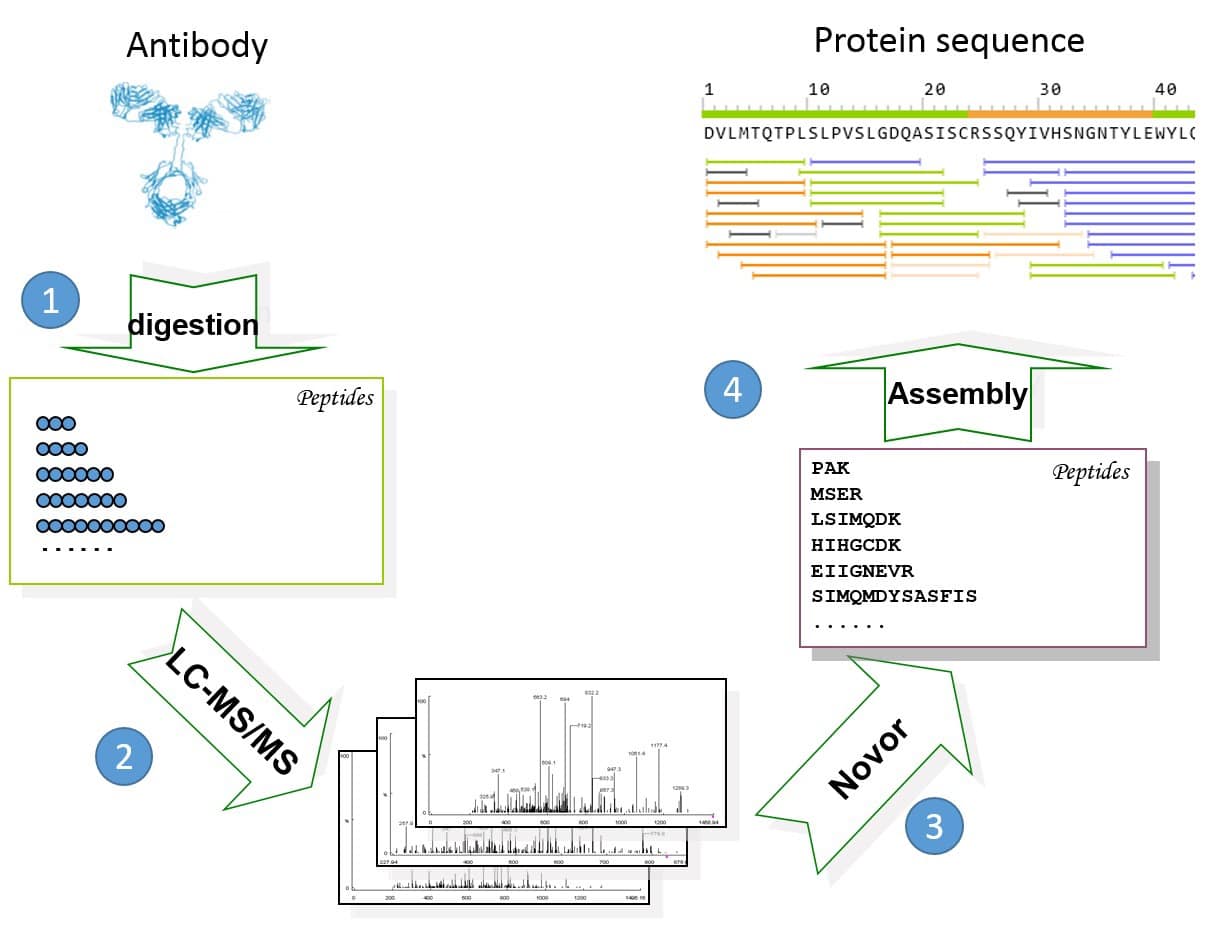
Mass Spectrometry-Based De Novo Sequencing of Monoclonal Antibodies Using Multiple Proteases and a Dual Fragmentation Scheme | Journal of Proteome Research
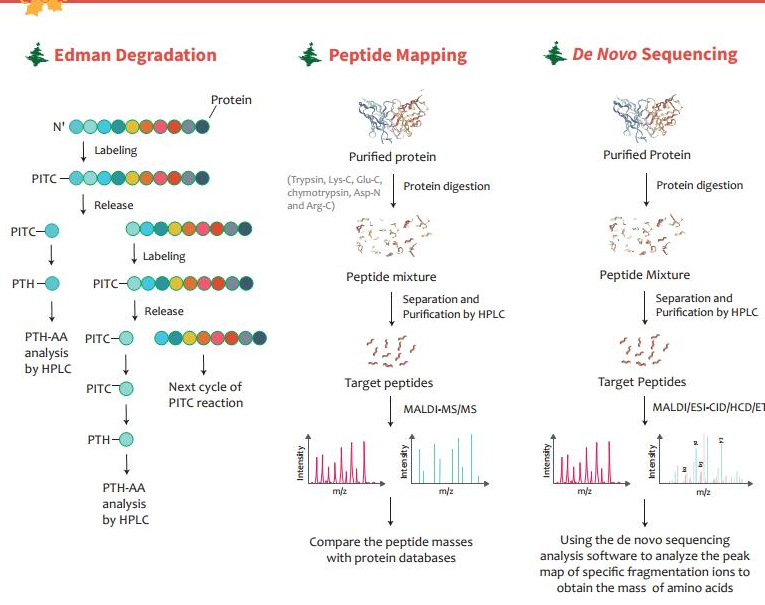
Creative Proteomics on X: "Chose suitable protein sequencing methods for your project This infographic introduces three protein sequencing methods: Edman degradation, peptide mapping, and de novo protein sequencing. https://t.co/kmI8xFuhlW https://t.co ...

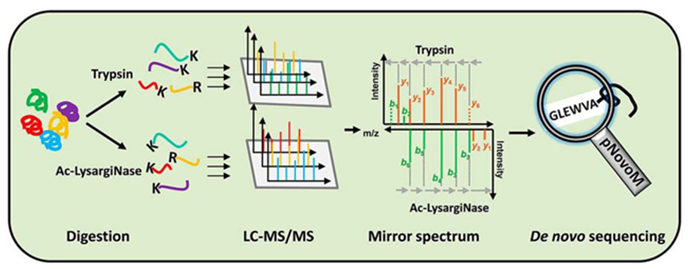
Precision De Novo Peptide Sequencing Using Mirror Proteases of Ac-LysargiNase and Trypsin for Large-scale Proteomics - ScienceDirect
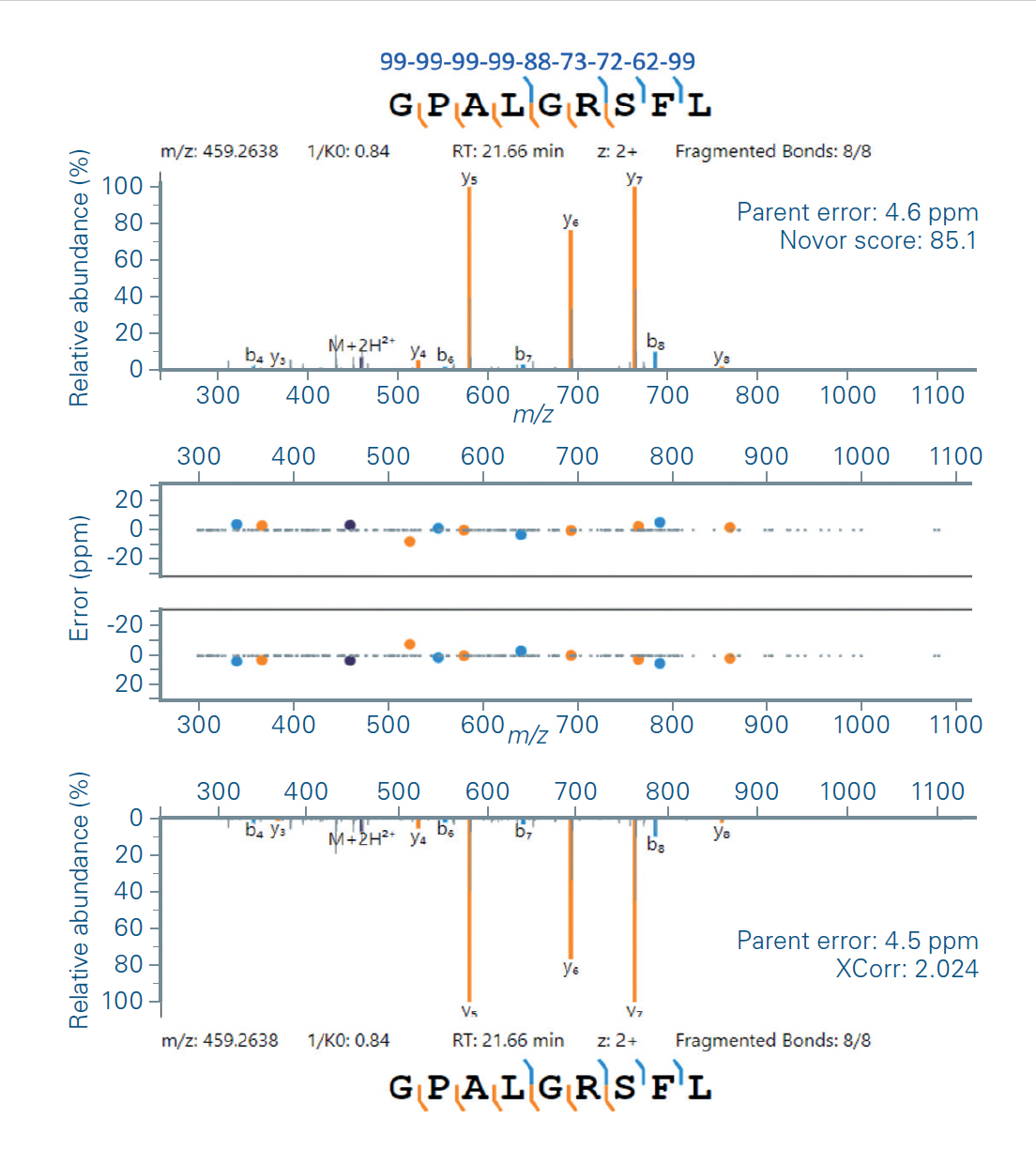
Bruker Launches de novo Sequencing for Immunopeptidomics, Library-Free dia-PASEF, Mass Dynamics Knowledge Visualization | Bruker
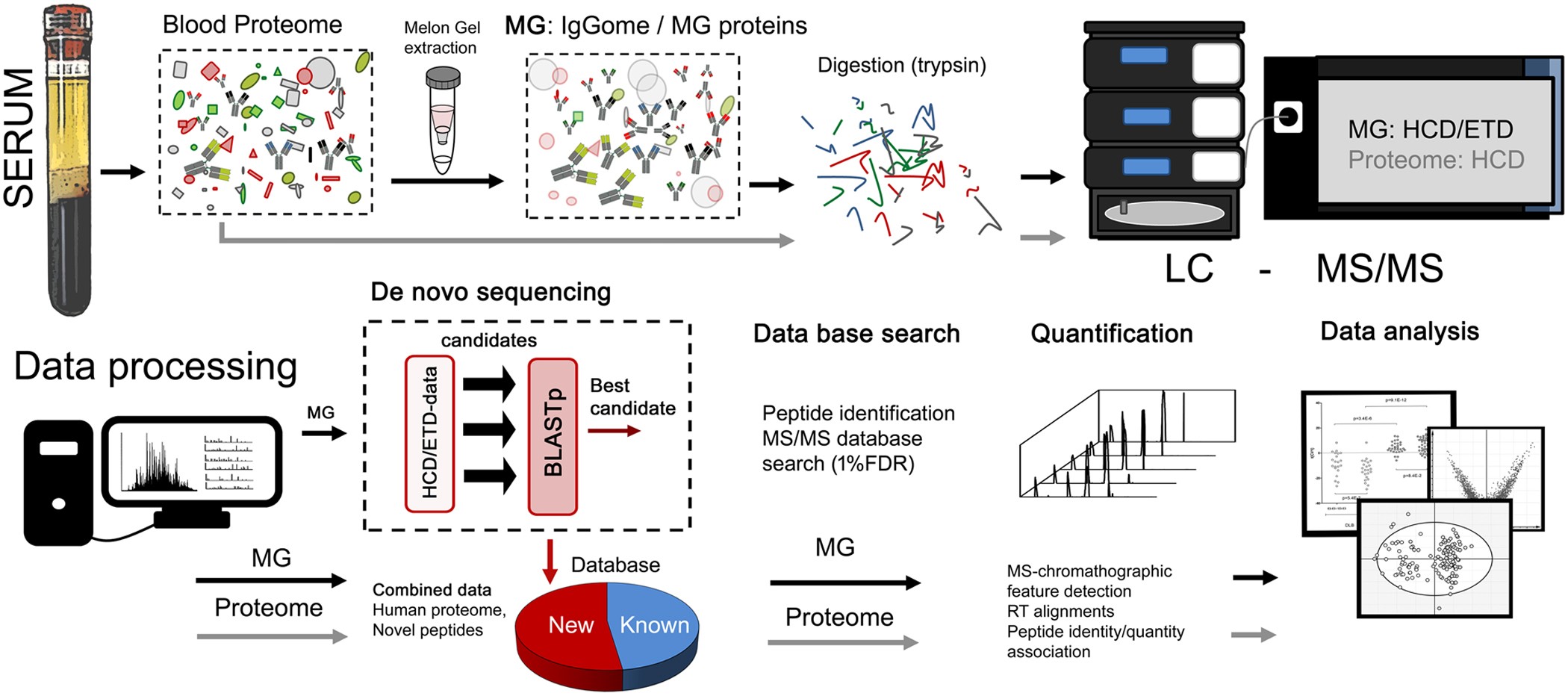
SpotLight Proteomics: uncovering the hidden blood proteome improves diagnostic power of proteomics | Scientific Reports